Protein Microarrays: A Powerful Tool for the Study of Autoantibodies
Autoimmune diseases are characterized by complex and heterogeneous antibody responses targeting a wide range of self-antigens. As highlighted in previous studies, these responses are often distributed across numerous targets, with individual reactivities that may be low in intensity yet biologically significant when considered collectively. This complexity requires analytical approaches capable of capturing the full breadth of immune reactivity rather than focusing on predefined panels of antigens.
Protein microarrays have emerged as a powerful technology for large-scale immune profiling, enabling the simultaneous interrogation of thousands of antigens in a single experiment. This approach provides a comprehensive view of antibody repertoires, supporting applications such as biomarker discovery, disease stratification, and the identification of immune signatures in oncology, autoimmunity, and infectious diseases. By capturing both expected and unexpected reactivities, protein arrays offer an unbiased perspective on complex immune responses.
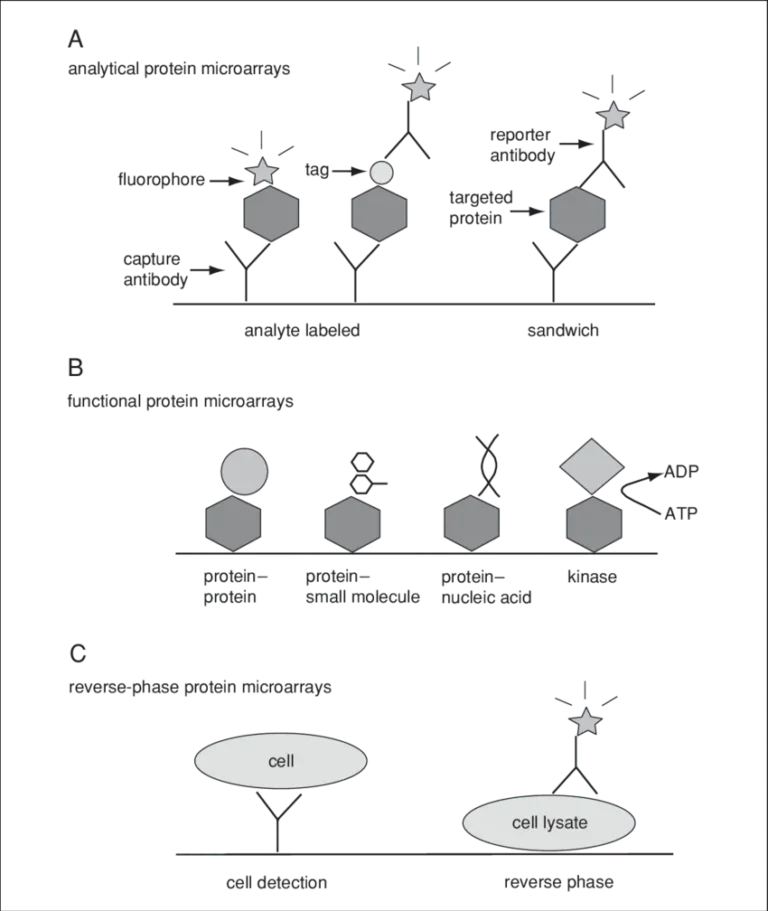
Figure 1: Three categories of protein microarrays.
(A) Analytical protein microarrays are mostly represented by antibody arrays and focus on protein detection. In this class of microarrays, targeted proteins can be detected either by direct labeling or using a reporter antibody in sandwich assay format.
(B) Functional protein microarrays have broad applications in studying protein interactions, including protein binding and enzyme-substrate reactions.
(C) Reverse-phase protein microarrays provide a different array format by immobilizing many different lysate samples on the same chip.
Exploring Autoimmunity: From Targeted Assays to Proteome-Wide Profiling
Let’s take a closer look at functionalized protein microarrays.
On these platforms, proteins are immobilized in their full-length, three-dimensional conformation, retaining key post-translational modifications and native folding. This enables biologically relevant interactions to occur directly on the array surface. As a result, protein–protein and antibody–antigen interactions can be interrogated with a level of fidelity that is not achievable with denatured proteins or peptide-based arrays.
The combination of structural integrity and broad proteome coverage is particularly powerful for uncovering unexpected immune reactivities and reducing the risk of overlooking critical disease drivers.
Beyond immune profiling, these arrays also open the door to functional studies of enzymatic activity, such as kinase-driven phosphorylation, ubiquitylation, acetylation or S-nitrosylation enabling systematic mapping of signaling networks at an unprecedented scale.
Autoantibody responses in patients with CBL deficiency
Building on these advances in large-scale immune profiling, recent studies such as the Somatic deficiency of the human E3 ubiquitin ligase CBL in leukocytes impairs B cell but not T cell development and function published in Nature Immulology highlight the value of next-generation protein microarrays that preserve protein structure and functionality.
In this study, the HuProt™ proteome-wide functional protein microarray (CDI Labs) was used to perform global autoantibody profiling in the serum of patients with CBL deficiency.
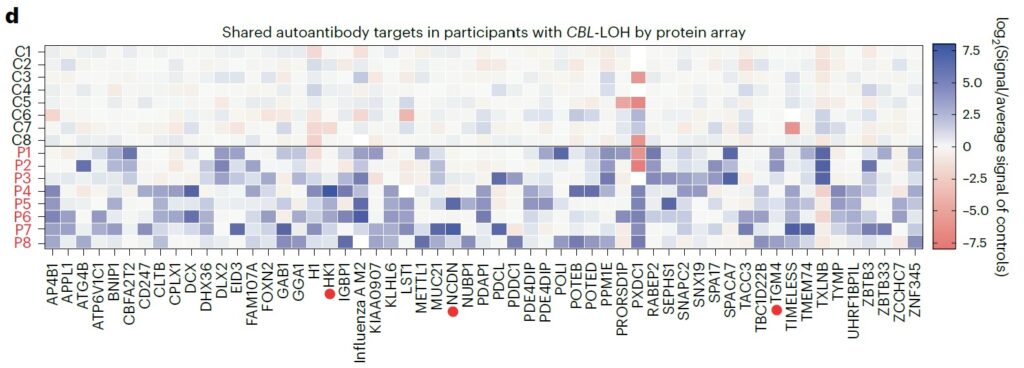
This approach enabled the identification of shared autoantigens across patients, while providing a quantitative and comparative assessment of antibody reactivity against thousands of full-length human proteins simultaneously. By capturing the complete spectrum of immune reactivity, the platform revealed a marked and widespread increase in autoantibody responses in patient samples compared to controls.
Notably, the analysis highlighted a broad expansion of autoreactivity rather than a restriction to a few dominant antigens, suggesting a global breakdown of immune tolerance. The data further showed that many of the identified targets were involved in key cellular pathways, including signaling and immune regulation, pointing to biologically relevant and potentially pathogenic interactions. This comprehensive profiling underscores the ability of proteome-wide arrays to uncover both shared and patient-specific immune signatures, providing deeper insight into disease mechanisms and identifying candidate biomarkers for further investigation.
Unparalleled Content, Reproducibility and Sensitivity for Autoantibody Discovery
The performance of these assays relies heavily on the quality of fluorescence detection achieved with the InnoScan range, recognized for its advanced imaging capabilities, low background noise, and consistent signal across the entire slide surface. With its high sensitivity and quantitative robustness, the InnoScan 1100 enables reliable detection of low-abundance autoantibodies, often biologically significant despite weak signal intensity, and supports accurate comparison across samples and cohorts. This level of performance is essential for identifying subtle yet meaningful differences in immune reactivity between individuals.

Combined with comprehensive, functional human proteome microarrays, the InnoScan 1100 enables enables reliable quantitative readout and high-throughput antibody validation. It goes beyond simply answering the question “Are autoantibodies present?”, it enables precise identification of what antibodies target at the proteome-wide level, with what intensity, and which shared patterns emerge across patients.
This level of precision is critical for translating complex immune profiling data into meaningful biological insights and actionable discoveries.
More than a reader, the InnoScan 1100 elevates protein microarrays from analytical tools to discovery platforms, opening new perspectives for clinical research, from early biomarker identification to a deeper understanding of immune-driven diseases at the proteome scale.
References
[1] Abel L, Kutschki S, Turewicz M, Eisenacher M, Stoutjesdijk J, Meyer HE, Woitalla D, May C. Autoimmune profiling with protein microarrays in clinical applications. Biochim Biophys Acta. 2014 May;1844(5):977-87. doi: 10.1016/j.bbapap.2014.02.023. Epub 2014 Mar 7. PMID: 24607371
[2] Sutandy FX, Qian J, Chen CS, Zhu H. Overview of protein microarrays. Curr Protoc Protein Sci. 2013 Apr;Chapter 27(1):Unit 27.1. doi: 10.1002/0471140864.ps2701s72. PMID: 23546620; PMCID: PMC3680110
[3] Ramachandran, N., Srivastava, S. and LaBaer, J. (2008), Applications of protein microarrays for biomarker discovery. Prot. Clin. Appl., 2: 1444-1459. https://doi.org/10.1002/prca.200800032
[4]Vatovec, T., Neehus, AL., Jackson, K.J.L. et al. Somatic deficiency of the human E3 ubiquitin ligase CBL in leukocytes impairs B cell but not T cell development and function. Nat Immunol 27, 308–322 (2026). https://doi.org/10.1038/s41590-025-02381-7
