μPharma: A microfluidic, AI-driven pharmacotyping platform for single-cell drug sensitivity prediction in leukemia
Huiqian Hu (1), Huanbin Zhao (2), Ping Lu (1), Teng Ma (1), Satoshi Yoshimura (2), Seth E. Karol (3), Ching-Hon Pui (3), David T. Teachey (4)
Jun J. Yang (2,3), Alphonsus H.C. Ng (1,5), and Yue Lu (1)
The μPharma platform (Hu et al., Med, 2026) demonstrates how high-resolution, multi-laser fluorescence scanning can serve as the analytical backbone of next-generation precision oncology. By combining digital microfluidics with quantitative single-cell fluorescence imaging, μPharma predicts leukemia drug sensitivity within four hours without direct drug exposure. The InnoQuant scanner enables multiplexed detection of total and phosphorylated proteins at 0.5 µm resolution, capturing not only protein expression levels but also spatial distribution
and morphological features at single-cell resolution. These high-content quantitative imaging data feed machine learning models that accurately predict therapeutic response while revealing intratumoral heterogeneity. This study illustrates how the Innopsys fluorescence scanning technology transforms microfluidic assays into AI-ready platforms, positioning InnoQuant as central enabler of rapid, scalable, and clinically actionable pharmacotyping in AI-based precision oncology.
Key findings: 📊 pBCL2 predicts venetoclax response (first time reported in T-ALL!) 📊 pLCK confirms dasatinib sensitivity 📊 Spatial protein distribution + cell morphology boost prediction accuracy
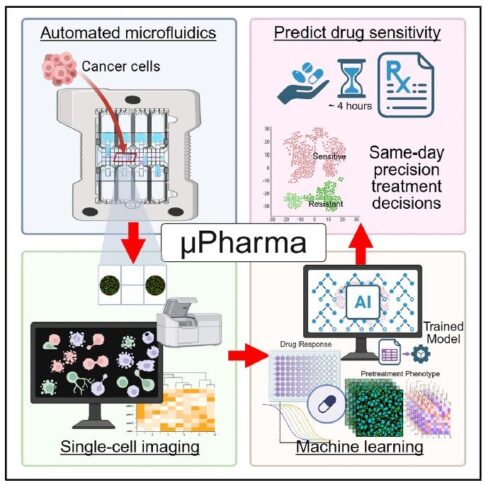
InnoQuant plays a core enabler role in this pharmacotyping platform:
✔ 0.5 µm/pixel resolution
✔ 4 lasers / 5 emission filters
✔ True multiplex (Actin, LCK, pLCK, BCL2, pBCL2)
✔ Minimal spectral bleed-through
✔ Single-cell spatial protein quantification
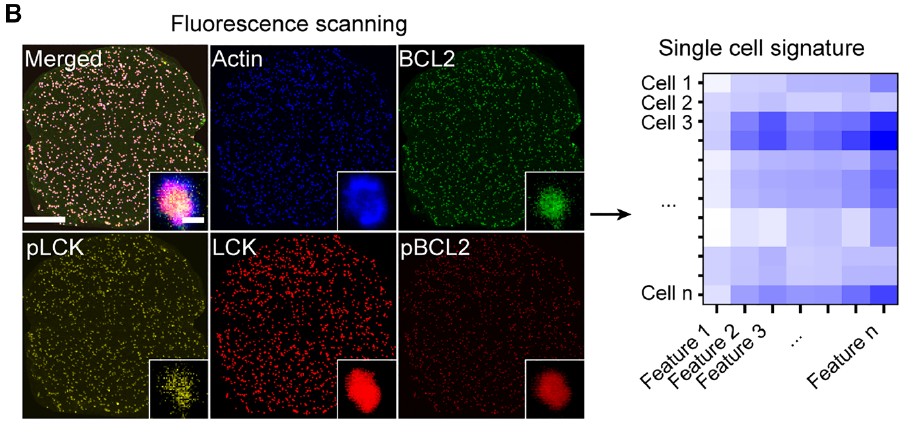
The point scanning process combined to PMT detection ensables a true quantitative extraction of:
- Protein intensity
- Phosphorylation status
- Morphological features
- Radial spatial distribution metrics
These features directly feed AI models for drug sensitivity prediction.
Our unique multi-laser fluorescence scanning technology is not just imaging but provides the quantitative engine powering AI-based precision oncology.
Access the article online: μPharma: A microfluidic, AI-driven pharmacotyping platform for single-cell drug sensitivity prediction in leukemia – ScienceDirect
1 Department of Molecular Pharmaceutics, University of Utah, Salt Lake City, UT 84112, USA
2 Department of Pharmacy and Pharmaceutical Sciences, St. Jude Children’s Research Hospital, Memphis, TN 38105, USA
3 Department of Oncology, St. Jude Children’s Research Hospital, Memphis, TN 38105, USA
4 Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA 19104, USA
5 Department of Biomedical Engineering, University of Utah, Salt Lake City, UT 84112, USA
